Luigi Ricciardi 1 , Rosa Mazzeo 2,*© , Angelo Raffaele Marcotrigiano 1 , Guglielmo Rainaldi 3 , Paolo Iovieno 4 , Vito Zonno 1 , Stefano Pavan 1© uye Concetta Lotti 2,*
- 1 Dhipatimendi revhu, Plant uye Food Sciences, Plant Genetics uye Breeding Unit University yeBari, Via Amendola 165/A, 70125 Bari, Italy; luigi.ricciardi@uniba.it (LR);angelo.marcotrigiano@uniba.it (ARM); vito.zonno@uniba.it (VZ); Stefano.pavan@uniba.it (SP)
- 2 Dhipatimendi reSainzi reKurima, Chikafu uye Zvakatipoteredza, University of Foggia, Via Napoli 25, 71122 Foggia, Italy.
- 3 Dhipatimendi reBiosciences, Biotechnologies uye Biopharmaceuticals, University of Bari, Via Orabona 4, 70125 Bari, Italy; guglielmo.rainaldi@uniba.it
- 4 Department of Energy Technologies, Bioenergy, Biorefinery and Green Chemistry Division, ENEA Trisaia Research Center, SS 106 Ionica, km 419+500, 75026 Rotondella (MT), Italy; paolo.iovieno@enea.it
* Correspondence: rosa.mazzeo@unifg.it (RM); concetta.lotti@unifg.it (CL)
Abstract:
Onion (Allium cepa L.) chirimwa chemuriwo chechipiri chakakosha pasi rose uye chinofarirwa nevakawanda nekuda kwehutano hwacho. Pasinei nekukosha kwayo kwehupfumi uye kukosha kwayo sechikafu chinoshanda, hanyanisi haina kuongororwa zvakanaka maererano nekusiyana kwayo kwemajini. Pano, takaongorora genetic variation mu "Acquaviva red onion" (ARO), landrace ine nhoroondo yekare yekurima mutaundi diki mudunhu reBari (Apulia, Southern Italy). Seti ye11 microsatellite marker yakashandiswa kuongorora kusiyanisa kwemajini muunganidzwa wegermplasm unosanganisira gumi nenhatu ARO vanhu uye matatu akajairika marudzi ekutengesa. Kuongororwa kwemajini maitiro ane parametric uye isiri-parametric nzira dzakaratidza kuti iyo ARO inomiririra yakanyatsotsanangurwa gene dziva, rakasiyana zvakajeka kubva kuTropea neMontoro landraces iyo inowanzokanganisa. Kuitira kupa tsananguro yemagirobhu, anowanzo kushandiswa kutsva, soluble solid content uye pungency zvakaongororwa, zvichiratidza kutapira kwepamusoro muARO maererano nembiri dzataurwa pamusoro apa. Pakazere, chidzidzo chazvino chinobatsira mune ramangwana kusimudzira kweARO, iyo inogona kusimudzirwa kuburikidza nemhando yemazita ayo anogona kubatsira kuderedza hutsotsi hwekutengesa nekuvandudza mari yevaridzi vadiki.
ziviso
The Allium genus inosanganisira dzinenge 750 marudzi [1], pakati peiyo hanyanisi (Allium cepa L., 2n = 2x = 16) ndeimwe yezvakanyanya kupararira. A. cepa ine biennial cycle uye kudarika maitiro ekubereka. Mazuva ano, kugadzirwa kwehanyanisi pasi rose (97.9 Mt) inoita kuti ive yechipiri yakakosha chirimwa chemuriwo mushure mematomatisi [2]. Kubva kare, magirobhu ehanyanisi ave achishandiswa zvese sechikafu uye mumishonga yevanhu. Chokwadi, vaIjipita vekare vakatotaura akati wandei maitiro ekurapa anoenderana nekushandiswa kwegariki nehanyanisi munhokwe yezvokurapa ye1550 BC, iyo Codex Ebers [3].
Muriwo uyu unoshanda zvakasiyana-siyana uye une hutano unodyiwa wakabikwa, mutsva, kana sechigadzirwa, uye unoshandiswa kusimudzira kuravira kwendiro dzakawanda. Ongororo dzinoverengeka dzechangobva kuitika dzinoti kunwa hanyanisi kunogona kuderedza njodzi yezvirwere zvemoyo [4,5], kufutisa [6], chirwere cheshuga [7], uye marudzi akasiyana ekenza [8-10]. Hutano hwehutano hwehanyanisi hunowanzonzi hune huwandu hwepamusoro hwemakirasi maviri ezve nutraceutical compounds: flavonoids uye alk(en)yl cysteine sulphoxides (ACSOs). Kirasi yekutanga inosanganisira flavonols uye anthocyanins. Quercetin ndiyo inonyanya kuoneka flavonol, inozivikanwa neayo yakasimba antioxidant uye anti-inflammatory properties mumahara radical scavenging uye transition metal ions kusunga. [11]; nepo anthocyanins inopa tsvuku/pepuru ruvara kune mamwe marudzi ehanyanisi. Kana ari maACSO, akanyanya kuwanda isoalliin [(+)-trans-S-1-propenyl-L-cysteine sulfoxide] [12], isina-kuvhuvhuta uye isina-proteinogenic sulfur amino acid yakachengetwa mumaseru, iyo inokonzera nenzira isina kunanga kunhuhwirira uye kuravira kwehanyanisi. [13]. Pamusoro pekuvhiringidzwa kwenyama, isoalliin inotsemurwa ne enzyme alliinase kuti ibudise mutsara wemakemikari anopisa (pyruvate, ammonia, thiosulphonates uye propanethial S-oxide) inokonzera kubvarura uye kukonzera kunhuhwirira kusingafadzi (pungency) [14]. Iyo onion pungency inowanzoyerwa sehuwandu, pagiramu rehuremu hutsva, hwepyruvic acid inogadzirwa nehydrolysis. [15, 16].
Munyika dzeMediterranean basin, inokurudzirwa seimwe yechipiri siyana nzvimbo dze A. cepa [17, 18], magirobhu ehanyanisi anoratidza kusiyana kwakasiyana-siyana kwechimiro, kukura, ruvara, chinhu chakaoma, uye pungency [19-22]. Uyezve, kusarufa-based fertilization, agronomic maitiro, mhando yevhu, mamiriro ekunze, uye genotype yezvirimwa kana landraces zvinogona kukanganisa girobhu mhando nekupa yakasarudzika organoleptic uye hutano hunhu. [23-27]. MuItaly, zvisinei nehupamhi hwehanyanisi huripo, mhando shoma dzehanyanisi dzinowanzo itwa zvidzidzo zvesainzi uye dzinozivikanwa nemazvo. [28, 29].
Kunyatsoita genetic uye phenotypic hunhu hweagro-biodiversity yakakosha kuvimbisa kuchengetedzwa kwakakodzera kwechirimwa genetic zviwanikwa uye kukurudzira kushandiswa kweiyo genotypes mucheni yekukosha. [30-32]. Yakareruka kutevedzana kudzokorora (SSR) mamaki agara achisarudzwa pamepu [33-35], DNA fingerprinting uye cultivar rusarura [36-38], uye fungidziro yakavimbika yekusiyana kwemajini mukati uye pakati penyika [39-42], sezvo iwo ari locus chaiwo, akawanda-allelic, codominantly nhaka, yakanyanya kudhindwa, uye akakodzera otomatiki genotyping.
Muchidzidzo chemazuva ano, takaisa pfungwa dzedu kune imwe nzvimbo yeApulian landrace, "Acquaviva red onion" (ARO), inorimwa maererano nemitoo yekurima mune imwe nzvimbo duku yeguta reAcquaviva delle Fonti, mudunhu reBari. (Apulia, Southern Italy). Ma bulbs eiyi landrace akakura uye akatsetseka uye ane ruvara rutsvuku uye anonyanya kushandiswa mumabikirwo emunharaunda. Kunyangwe iyo ARO yakawana "Slow Chikafu Presidium" chiratidzo chemhando, kugadzirwa kwayo kunogona kusimudzirwa nekudzivirirwa neEuropean Union mhando mamaki senge akadzivirirwa geographical chiratidzo (PGI) uye akadzivirirwa kudomwa kwekwakabva (POD), sezvo izvi zvichigona kubatsira kuderedza hutsotsi hwekutengesa nekuvandudza mihoro yevarimi vadiki. Pano, SSR mamorekuru emakaka akashandiswa sezvishandiso zvine simba kuongorora genetic variation pakati pehuwandu hweARO uye kusarura iyi landrace kubva kune dzimwe mbiri dzeSouthern Italy red onion landraces. Uyezve, isu takafungidzira pungency uye soluble yakasimba yemukati kuitira kuti tiongorore ARO flavour inoenderana nekudiwa kwemusika.
Results
Kugadzwa kweAcquaviva Red Onion Germplasm Collection uye Morphological Characterization
Mbeu dzine vanhu gumi nenhatu dzeARO landrace, dzakapihwa nevarimi muchirongwa cheBiodiverSO Apulia Region project dzakashandiswa kutanga kuunganidzwa kwegermplasm yeARO.
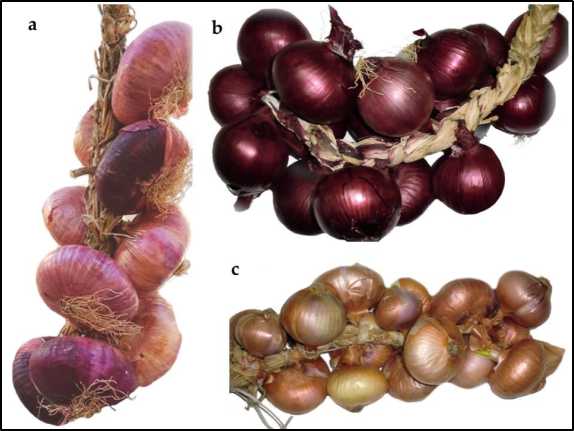
Morphological descriptors, zvine chekuita negirobhu, ganda, uye nyama zvakaunganidzwa paARO germplasm uye pamatatu ehanyanisi landrace, maviri ari e "Tropea red onion" (TRO) landrace uye imwe ye "Montoro copper onion" (MCO) landrace (Mufananidzo. 1). Ese maARO bulbs aive akati sandarara uye aionekwa neganda dzvuku rekunze nenyama ine mumvuri wakasiyana wehutsvuku. Kusiyana neizvi, nyama yeTRO bulbs yaive yakatsvuka zvakazara, nepo nyama yeMCO bulbs yaive isina pigmented (Tafura S1). Ongororo yeBiochemical inobvumidzwa kuongorora iyo yakasimba soluble yemukati uye pungency. Sezvakataurwa muTable 1, huremu hwezvakasimba zvinonyungudika zvema bulbs muhuwandu hweARO hwaive 7.60, uye kubva pa6.00 (ARO12) kusvika pa9.50° Brix (ARO11 neARO13). Ukoshi uhwu hwaive hwepamusoro pane hunofungidzirwa hweTRO neMCO landraces (4.25 uye 6.00 ° Brix, zvichiteerana).

Tafura 1. Solid Soluble Content uye Pungency Values Yakaongororwa mu "Acquaviva Red Onion" (ARO), "Tropea Red Onion" (TRO), uye "Montoro Copper Onion" (MCO) Populations *.
| CODE | Soluble Solid Content (Brix) | Pungency (pmolg-1 FW) | ||
| Zvinoreva | CV y (%) | Zvinoreva | CV y (%) | |
| ARO1 | 6.25 D * | 5.65 | 5.84 ab * | 23.78 |
| ARO2 | 7.25 DC | 4.87 | 6.51 kuti | 22.98 |
| ARO3 | 7.50 BCE | 9.42 | 5.28ab | 22.88 |
| ARO4 | 7.50 BCE | 0.00 | 6.97 kuti | 3.74 |
| ARO 5 | 7.50 BCE | 0.00 | 6.80 kuti | 9.68 |
| ARO6 | 6.25 D | 5.65 | 4.51ab | 39.18 |
| ARO7 | 7.25 DC | 4.87 | 5.25ab | 15.44 |
| ARO8 | 9.00 AB | 0.00 | 7.04 kuti | 3.49 |
| ARO9 | 8.25 ABC | 4.28 | 6.84 kuti | 0.15 |
| ARO10 | 7.00 DC | 0.00 | 5.94ab | 6.57 |
| ARO11 | 9.50 A | 7.44 | 5.54ab | 16.43 |
| ARO12 | 6.00 D | 0.00 | 4.91ab | 9.70 |
| ARO13 | 9.50 A | 7.44 | 6.63 kuti | 24.93 |
| MCO | 6.00 D | 0.00 | 4.18ab | 2.66 |
| TRO1 | 4.25 E | 8.31 | 2.80 b | 2.10 |
| TRO2 | 4.25 E | 8.31 | 4.28ab | 4.79 |
* Zvinoreva nemabhii mamwe chete muhukuru kana madiki hazvina kusiyana pa 0.01P kana 0.05P, zvichiteerana (SNK's Test). y Coefficient of variation.
Iko kukosha kweARO pungency, yakaongororwa nenzira yepyruvic acid content, yaiva 6.00, yakabva ku4.51 pmol g.-1 FW (ARO6) kusvika 7.04 (ARO8). Ukoshi uhwu hwaive hwepamusoro kupfuura hunofungidzirwa muTRO neMCO landraces (3.54 pmol g-1 FW uye 4.18 pmol g-1 FW, zvichiteerana).
SSR Polymorphism uye Genetic Relationships pakati peAccessions
Muchidzidzo chazvino, gumi nerimwe kubva pamakumi matatu nenomwe akaedzwa SSR primer musanganiswa akapa imwechete-locus polymorphisms, kureva, kuburitsa akawanda maviri ekuwedzera zvigadzirwa mumunhu mumwe. Pakazere, 11 alleles dzakaonekwa muvanhu mazana matatu nemakumi maviri vane nhamba yealleles per locus kubva pa37 (ACM55 uye ACM 320) kusvika 2 (ACM147) uye kukosha kwe504 alleles (Table 2). Muvanhu vega, nhamba ye alleles (Na) yakabva ku1.94 (ACM147 uye ACM504) kusvika ku5.38 (ACM132), nepo nhamba inoshanda yealleles (Ne) yakabva ku1.41 (ACM152) kusvika ku2.82 (ACM449). Kusiyana pakati peNa neNe tsika dzakakonzerwa nekuvapo kwe alleles ine yakaderera frequency muhuwandu hwevanhu uye predominance yezvishoma alleles. Iyo yakanyanya kucherechedzwa heterozygosity (Ho) kukosha kwakaratidzwa kune ACM138 uye ACM449 (0.62), nepo yakaderera yakabatana neACM152 (0.25). Inotarisirwa heterozygosity (Iye), iyo inofananidzwa netarisiro yezvinyorwa muhuwandu hwepanmictic, yakabva ku0.37 (ACM504) kusvika ku0.61 (ACM132, ACM138, uye ACM449). Iyo Wright's fixation index (Fis), yakaratidza hunhu huri pedyo ne zero (avhareji 0.05) kune zvese zvinomaka, zvichiratidza hunhu hwakafanana pakati peinocherechedzwa uye inotarisirwa heterozygosity mwero, sezvaitarisirwa kune inopfuura mhando yemhando. Kubudirira kwemunhu mumwechete SSR marker mu genetic fingerprinting yakafungidzirwa ne polymorphic information content (PIC) index, ine kukosha kwe0.48 uye yakabva ku0.33 (ACM504) kusvika ku0.67 (ACM132). Imwe indekisi inoshanda, iyo Shannon's Information Index (I) yakaratidza kukosha kwe0.84, uye yaifungidzirwa kukosha kubva pa0.45 (ACM152) kusvika 1.20 (ACM132).
Tafura 2. Polymorphism Zvimiro zve11 SSR Makaki Akashandiswa Kufungidzira Genetic Diversity muARO, TRO, uye MCO Populations. Nhamba Yese yeAlleles (Na), Band Size Range, uye Polymorphic Information Content (PIC) Index Tarisa kune Yese Seti ye320 Individuals Genotyped muchidzidzo chino. Nhamba yeAlleles (Na), nhamba yeAlleles Anobudirira (Ne), Akacherechedza Heterozygosity (Ho), Inotarisirwa Heterozygosity (Iye), Fixation Index (Fis), uye Shannon's Information Index (I) inoreva Kurevesa Values Yakaverengerwa kubva muvanhu gumi nevatanhatu, Imwe neimwe Inoumbwa ne16 Vanhu.
| Locus. | Total Na | Size Range (bp) | PIC | Zvinoreva | |||||
| Na | Ne | Ho | He | I | Fis | ||||
| ACM91 | 4 | 189-205 | 0.40 | 2.63 | 1.72 | 0.38 | 0.39 | 0.66 | 0.04 |
| ACM101 | 4 | 229-241 | 0.52 | 2.94 | 2.37 | 0.53 | 0.56 | 0.92 | 0.06 |
| ACM132 | 11 | 186-248 | 0.67 | 5.38 | 2.78 | 0.55 | 0.61 | 1.20 | 0.09 |
| ACM138 | 5 | 242-272 | 0.66 | 3.69 | 2.82 | 0.62 | 0.61 | 1.09 | -0.02 |
| ACM147 | 2 | 264-266 | 0.37 | 1.94 | 1.83 | 0.44 | 0.44 | 0.62 | -0.01 |
| ACM152 | 4 | 228-244 | 0.25 | 2.38 | 1.41 | 0.25 | 0.27 | 0.45 | 0.07 |
| ACM235 | 4 | 286-298 | 0.41 | 2.81 | 1.77 | 0.44 | 0.41 | 0.72 | -0.06 |
| ACM446 | 6 | 108-120 | 0.56 | 3.50 | 2.48 | 0.49 | 0.58 | 1.01 | 0.16 |
| ACM449 | 8 | 120-140 | 0.66 | 4.88 | 2.82 | 0.62 | 0.61 | 1.18 | -0.03 |
| ACM463 | 5 | 202-210 | 0.47 | 3.38 | 1.95 | 0.46 | 0.48 | 0.83 | 0.05 |
| ACM504 | 2 | 188-192 | 0.33 | 1.94 | 1.64 | 0.30 | 0.37 | 0.54 | 0.20 |
| Zvinoreva | 5 | 0.48 | 3.22 | 2.15 | 0.46 | 0.48 | 0.84 | 0.05 |
Pakati pehuwandu hwevanhu, ARO3, ARO6, ARO8, ARO10, TRO1, uye MCO yakaratidza huwandu hwepamusoro hwekusiyana kwemajini (Ho> 0.5), asi kusiyana kwakaderera kwakaonekwa muhuwandu hwevanhu ARO7 (Ho = 0.27) (Supplementary Table S2). Pakazara, zvese zvakabatanidzwa zvakaratidza Fis kukosha pedyo ne zero (Fis zvinoreva kukosha = 0.054), sezvinotarisirwa pasi pemamiriro ezvinhu ekubatana.
Kuongororwa kweMolecular Variance uye Genetic Structure
Hierarchical partitioning ye genetic variation pakati uye mukati mevanhu yakaverengerwa neAMOVA. Zvigumisiro zvakaratidza chikamu chikuru chekusiyana kwemajini mukati mevanhu (87%). Kusiyana pakati pevanhu, 13%, kwakakosha zvikuru (P <0.001) (Tafura 3). Pairwise tsika dzeFpt parameter, inofananidzwa neWright's Fst fixation index, kubva pa0.002 (ARO2/ARO10) kusvika 0.468 (ARO7/TRO2), yaikosha (P <0.05), kunze kwekufananidza zvipfumbamwe paviri (Supplementary Table S3).
Tafura 3. Analysis of Molecular Variance of 320 Genotypes kubva ku16 Populations of allium cepa L.
| mabviro | df | Sum yezvikwere | Variance Estimation | Kusiyana (%) | Fpt | P |
| Pakati pehuwandu hwevanhu | 15 | 458.63 | 1.16 | 13% | ||
| Mukati mehuwandu hwevanhu | 304 | 2272.99 | 7.50 | 87% | 0.134 | 0.001 |
| muunganidzwa | 319 | 2731.62 | 8.66 |
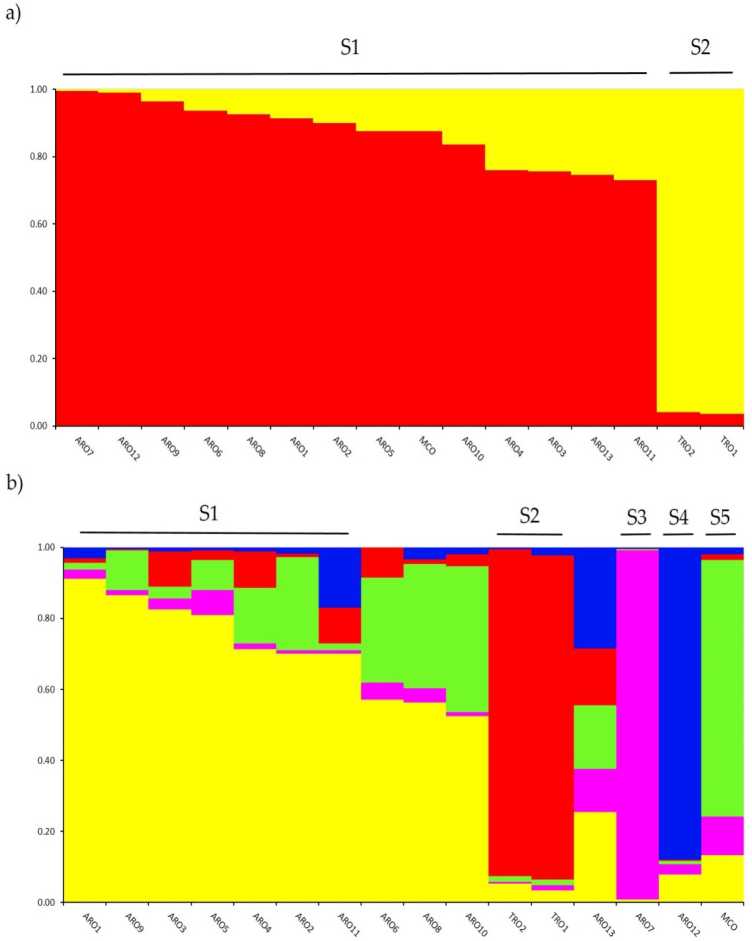
Kuongorora kwemajini chimiro mu A. cepa muunganidzwa we genotyped muchidzidzo ichi wakaitwa nenzira ye admixture model-based clustering analysis yakaitwa musoftware STRUCTURE. Iyo Evanno AK nzira yakaratidza kupatsanurwa mumasumbu maviri (K = 2) seyakanyanya kudzidzisa kune yedu. dhatabheti,na the Inotevera yepamusoro peak uye k = 5 (sopplementaiv Rgure S1). A zve K = 2, ahpopmatambudziko were assigned kuti onand thef masumbu maviri ne a rnernbertoip coefficient (q)> 0.7. Sezvo shown mukati figiri 2a, boka rekutanga (rakanzi S1) raisanganisira MCO uye vanhu vose veARO, nepo S2 cluster yakaronga mapoka maviri eTRO. PaK = 5, ichipa tsananguro yakadzama yedataset (Mufananidzo 2b), 75% yezviwanikwa zvakapihwa kune rimwe remasumbu mashanu. Kupatsanurwa pakati peARO (S1) neTRO (S2) kwakasimbiswa, kunyangwe mamwe maARO akasanganiswa (q <0.7) kana kuiswa akaparadzana mumasumbu maviri matsva S3 neS4 (ARO7 neARO12, zvichiteerana). Sezvineiwo, iyo MCO yekutengesa mhando yakagadzira rakasiyana sumbu (S5) rakaparadzaniswa neApulian red onion.
Genetic Relationships pakati peVanhu
SSR polymorphism inobvumirwa kutora dendrogram ye genetic diversity uye migumisiro ye phylogenetic analysis inoratidzwa mumufananidzo. 3a. Pano, iyo germplasm collection yakakamurwa mumapoka mashanu akatsigirwa zvakasimba nebootstrap values. Vanhu veARO7 neARO12 vakabva vaparadzaniswa kubva kuvanhu vakanga vasara ndokuumba mapoka maviri akasiyana. Boka rechitatu raisanganisira vanhu vaviri vekutengesa veTRO, ukuwo nzvimbo yechina yakapatsanura MCO kubva kugumi nerimwe vanhu veARO. Ukama hweGenetic huri kuitika pakati pehuwandu hwakaongororwa zvakare nenzira yeprincipal coordinate analysis (PCoA) (Mufananidzo 3b) Sezvambotaurwa, huwandu hweARO hwakaiswa mumapoka akasimba, kunze kweARO12 neARO7, iyo yakaonekwa munzvimbo dzakasarudzika muPCoA zano. Iwo maviri maTRO uye MCO huwandu hwevanhu vakapararira muzasi-kurudyi pani yechirongwa.
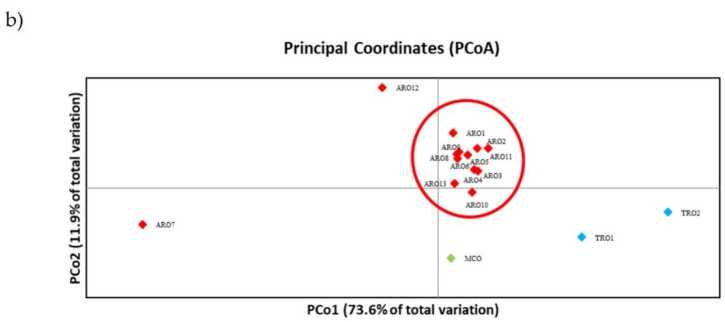
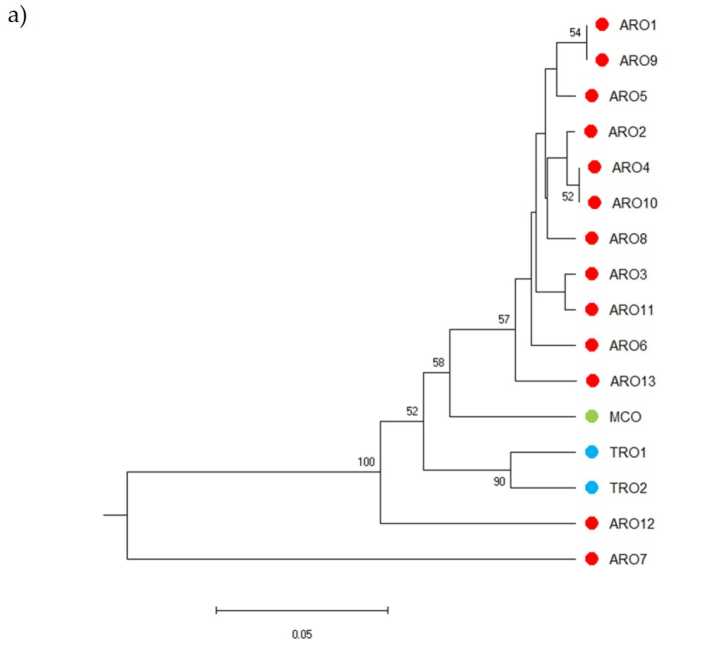
Mufananidzo 3. Genetic kusiyana pakati pe16 A. cepa vanhu vanoonekwa muchidzidzo ichi, zvichibva pane yavo SSR mbiri. (a) UPGMA dendrogram ye genetic kure. Bootstrap tsigiro tsika> 50 inoratidzwa pamusoro pemanodhi anoenderana; (b) principal component analysis (PCoA). Iro sumbu rakatenderedzwa mutsvuku rakanyatsoenderana neboka rakagadzirwa ne phylogenetic analysis uye rakagadzirwa ne 11 ARO accessions.
hurukuro
Mukati mehuwandu hukuru hwezvipenyu zvakasiyana-siyana zvinorimwa kuMaodzanyemba kweItaly, onion landraces inomiririra zvigadzirwa zvinoda kuchengetedzwa kubva panjodzi yekukukurwa kwemajini uye kutyisidzirwa kwekutsiviwa nembeu dzemazuva ano. Muhurongwa hwepurojekiti yedunhu BiodiverSO, yakanangana nekuunganidza, kuratidza, kusimudzira, nekuchengetedza zviwanikwa zvedunhu reApulia zvine hukama zvakanyanya nenhaka yenzvimbo, takagadzira muunganidzwa wembeu wevanhu gumi nevatatu veARO landrace. Takashuma kuongororwa kwekutanga kwekusiyana kweARO maererano neDNA polymorphisms uye maviri biochemical parameters, soluble solid uye pyruvic acid zviri mukati, zvine chekuita nekunaka hunhu uye kukosha kwekugamuchirwa kwezvitsva zvisina kubikwa zvigadzirwa. Pamusoro pezvo, data pamusoro peARO landrace yakaenzaniswa neiya yakaunganidzwa pane mamwe maviri ane pigmented onion landraces ayo aiwanzokanganisa nawo.
Ongororo yeBiochemical yakaratidza kutapira kwehuwandu hwegumi nenhatu ARO, ine hukama nepamusoro soluble solid yemukati uye yepakati pungency, zvinoenderana neinotapira indasitiri nhungamiro. [31]. MaARO bulbs aitapira kupfuura aya eTRO neMCO landraces, uye airatidza pungency yakakwira zvishoma. Zvisinei, kutapira muhanyanisi kunokonzerwa nekuenzana pakati peshuga uye pungency, saka chimiro ichi chinogona kubatsira kutsigira kusarudzwa kwema genotypes ekukosha, kazhinji kunoitwa nevarimi chete zvichienderana ne morphology.
Zvicherechedzo zveSSR zvakasimbiswa kuva chishandiso chinobatsira pakusarura genotypes, kunyangwe zvakaunganidzwa mukati menzvimbo yakamanikana inokura sedhorobha reAcquaviva delle Fonti. Iwo ma marker akasarudzwa airatidza nhamba yepamusoro ye alleles pane ma marker akambotaurwa na [43] uye [44], asi yakaderera pane mamaki akataurwa na [45]. Uyezve, 50% yezvicherechedzo zvedu zvakaratidza PIC index hukuru kupfuura 0.5, zvichiratidza kuti yakakodzera kusarura vanhu vari muunganidzwa, sezvakakurudzirwa na. [46]. Ongororo yekusiyana-siyana mukati mehuwandu yakaratidza hunhu hwakafanana pakati peHo naHe, zvichikonzera kuderera kweFis values. Izvi zvinopindirana nekunze-kuyambuka chimiro che A. cepa, iyo inotambura zvakanyanya ne inbreeding depression [47]. Iyo yakazara Fis kukosha kwakaverengerwa muhuwandu hwehanyanisi hwakatariswa muchidzidzo ichi (0.054) yaive yakaderera pane iya yakambotaurwa na. [45] (0.22) uye yakada kufanana neyakawanikwa na [31] (0.08) uye [48] (0.00) akaongorora kusiyana kwemajini mumijaho yehanyanisi kubva kuchamhembe kwakadziva kumadokero kweSpain neNiger, zvichiteerana. Zvinocherechedzwa mazinga eheterozygosity muhuwandu hweARO anosimbisa pfungwa yekuti Apulia inomiririra nzvimbo yakasiyana-siyana yemarudzi akawanda ehorticultural. [32, 42, 49-51].
AMOVA yakaratidza kuti mutsauko wakawanda wemamorekuru muunganidzwa wakadhindwa muchidzidzo ichi uri mukati mehuwandu. Nekudaro, yakakosha mutsauko wemajini pakati pehuwandu (FPT values) zvakaratidza kuitika kwe genetic stratification. Kutaura zvazviri, kunyange zvazvo migumisiro yedu yakaratidza kuvapo kwemajini akafanana muhuwandu hwevanhu veARO, kuumba boka rinonyatsotsanangurwa, ARO7 uye ARO12 vanhu vakaratidza maitiro akasiyana zvakajeka. Mhedzisiro iyi inogona kunge yakakonzerwa nekwakasiyana kwakabva mbeu dzakashandiswa nevarimi vaviri uko kwakatorwa vanhu. Uyezve, zvichibva pamhedzisiro yakawanikwa, iyo ARO landrace inogona kutariswa zvakajeka padanho re genetic kubva kuTRO neMCO landraces. Muongororo yakaitwa munguva pfupi yapfuura, [29] yakaongorora kusiyana kwemajini emarudzi akawanda eehanyanisi eItaly anosanganisira "Acquaviva," "Tropea," uye "Montoro." Kunyange zvazvo vanyori vakashandisa zviratidzo zveSNP kuti vaongorore kusiyana kwemajini ekuunganidza kwehanyanisi yakawanda, genotyping haina kukwanisa kusarura "Acquaviva" kubva ku "Tropea" uye "Montoro" anyanisi. Pamwe, kusawirirana uku kunokonzerwa nekushomeka kwePIC kukosha kwakawanikwa (0.292), zvichiratidza ruzivo rune mwero rweiyo loci iri kuongororwa sezvakataurwa na. [29]. Uyezve, kuitira kuti tiongorore kuvapo kwe-sub-structure muchikwata chavo cheItaly, zvingave zviri nani kuongorora maItaly genotypes zvakasiyana kubva kune mamwe ose ekuunganidza. Pamwe ingadai yakabvumira kufungidzira maitiro ekusiyana kwemajini kwakabatana neiyo geographic stratification kana hunhu pasi pe empiric sarudzo.
Mukupedzisa, chidzidzo chazvino chinomiririra mushumo wakakwana pamusoro peanion landrace ine chekuita nenhaka yetsika nemagariro uye kukosha kwehupfumi kuvarimi. Zvigumisiro zvedu zvinoratidza kuti, kunze kwezvishoma zvishoma, ARO inoratidzirwa nejini yakanyatsotsanangurwa, iyo inofanirwa kuchengetedzwa kubva pangozi yekukanganiswa kwemajini. Naizvozvo, kugadzwa kwemuunganidzwa wemumiririri weiyi yakakosha sosi yekusiyana kwemajini kwave kwakakosha. Chekupedzisira, iyo genetic uye phenotypic characterization yeARO inogona kubatsira kuwana mhando mamakisi kubva kuEuropean Union.
Zvinhu uye Nzira
Kuunganidzwa kweGermplasm, Chimiro Chekudyara, uye Kutorwa kweDNA
Seti yegumi nenhatu vanhu veARO landrace vakawanikwa mukati mehurongwa hweApulia Dunhu chirongwa (BiodiverSO: https://www.biodiversitapuglia.it/), kuburikidza nenhevedzano yemamishinari akaitwa mu "Acquaviva delle Fonti", guta diki reApulian muProvince yeBari, Italy. Nzvimbo dzekuunganidza dzega dzega dzakaiswa mamepu kuburikidza neGeographic Information System (GIS) uye dzakataurwa muTafura 4. Pamusoro pezvo, vanhu vaviri kubva kuTRO landrace uye munhu mumwe kubva kuMCO landrace vakaverengerwa muchidzidzo chezvino uye vakashandiswa semareferenzi. Yese chirimwa chakarimwa mumamiriro ezvakatipoteredza papurazi rekuyedza "P Martucci" yeYunivhesiti yeBari (41 ° 1'22.08 ″ N, 16 ° 54'25.95 ″ E), pasi pekeji yekudzivirira kudzivirira kuyambuka pakati. uwandu hwevanhu uye kuvimbisa kufambiswa kwemukume kuburikidza nenhunzi (Lucilia Kesari). Idzo 16 vanhu vaizivikanwa hunhu hune hukama nehukuru hwebhubhu uye chimiro uye ganda nenyama ruvara (Table S1). Pamusoro pezvo, solid soluble content assay yakaitwa pachishandiswa ruoko-inobata refractometer uye pungency yakayerwa muonion juice samples ichiwedzera 2,4-dinitrophenyl hydrazine (0.125% v/v mu2N yeHCl) uye kuongorora kunyura pa420 nm, sezvakataurwa na [31]. Iyo Duncan's akawanda-renji bvunzo uye iyo SNK bvunzo yakaitwa kuti ione kuvepo kwekukosha kwakasiyana.
Tafura 4. Rondedzero Yehuwandu Hwakaunganidzwa uye Genotyped muChidzidzo Ichi. Kune Vese Vagari, Identification Code, Zita reNzvimbo, GPS Coordinate, uye Gene Bank Kuchengetedza Mbeu zvinoshumwa.
| kodhi | zita | GPS Inobatanidza | Gene Bank y |
| ARO1 | Cipolla rossa di Acquaviva | 40°54’21.708″ N 16°49’1.631” E | Di.SSPA |
| ARO2 | Cipolla rossa di Acquaviva | 40°53’14.28″ N 16°48’56.879” E | Di.SSPA |
| ARO3 | Cipolla rossa di Acquaviva | 40°54’11.304″ N 16°49’13.079” E | Di.SSPA |
| ARO4 | Cipolla rossa di Acquaviva | 40°54’3.348″ N 16°40’27.011” E | Di.SSPA |
| ARO5 | Cipolla rossa di Acquaviva | 40°51’59.76″ N 16°53’0.527” E | Di.SSPA |
| ARO6 | Cipolla rossa di Acquaviva | 40°52’48.72″ N 16°49’43.247” E | Di.SSPA |
| ARO7 | Cipolla rossa di Acquaviva | 40°53’13.47″ N 16°50’23.783” E | Di.SSPA |
| ARO8 | Cipolla rossa di Acquaviva | 40°53’18.816″ N 16°49’33.888” E | Di.SSPA |
| ARO9 | Cipolla rossa di Acquaviva | 40°54'51.372″ N 16°49'3.504" E | Di.SSPA |
| ARO10 | Cipolla rossa di Acquaviva | 40°54’1.188″ N 16°49’24.311” E | Di.SSPA |
| ARO11 | Cipolla rossa di Acquaviva | 40°52'49.8″ N 16°49'48.575" E | Di.SSPA |
| ARO12 | Cipolla rossa di Acquaviva | 40°52’38.892″ N 16°49’28.379” E | Di.SSPA |
| ARO13 | Cipolla rossa di Acquaviva | 40°53’21.768″ N 16°49’29.711” E | Di.SSPA |
| TRO1 | Cipolla rossa lunga di Tropea | - | Di.SSPA |
| TRO2 | Cipolla rossa tonda di Tropea | - | Di.SSPA |
| MCO | Cipolla ramata di Montoro | - | Di.SSPA |
| y Di.SSPA, Dhipatimendi reIvhu, Chidyarwa uye Chikafu Sayenzi, Yunivhesiti yeBari. |
Mashizha e20 genotypes pahuwandu akatorwa uye akachengetwa pa -80 °C kusvika ashandiswa. Kune marudzi ane polysaccharide-rich, se A. cepa, Matanho ekutanga ekubvisa polysaccharide akakosha kuti uwane yakanaka-mhando yeDNA, saka ekutanga anogeza muSTE buffer (0.25 M sucrose, 0.03 M Tris, 0.05 M EDTA) akaitwa sezvakatsanangurwa ne [52]. Yese DNA yakatorwa ichitevera nzira yeCTAB [53] uye pakupedzisira yakatariswa kunaka uye kutariswa neNano Drop 2000 UV-vis spectrophotometer (ThermoScientific, Waltham, MA, USA) uye 0.8% agarose gel electrophoresis.
SSR Analysis
16 EST-SSR primer musanganiswa wakagadzirwa ne [54] uye akamboedzwa muzvidzidzo zvemajini akasiyana-siyana na [43] uye [44] uye 21 genomic SSR [45-55] vakaongororwa kuti vaone kukodzera kwavo (Supplementary Table S4). Genotyping yakaitwa uchishandisa yehupfumi fluorescent tagging nzira umo iyo M13 muswe inowedzerwa kune yega kumberi SSR primer. [56]. PCR misanganiswa yakagadzirwa mu20 gL reaction ine: 50 ng yeDNA yakazara, 0.2 mM yedNTP musanganiswa, 1X yePCR reaction buffer, 0.8 U ye DreamTaq DNA polymerase (Thermo Scientific, Waltham, MA, USA), 0.16 gM ye reverse primer. , 0.032 gM yekutangira kumberi yakawedzerwa neM13 kutevedzana (5′-TGTAAAACGACGGCCAGT-3′), uye 0.08 gM yepasirese M13 primer yakanyorwa neFAM kana NED fluorescent dhayi (Sigma-Aldrich, St. Louis, MO, USA). Maitiro ePCR akaitwa muSimpliAmp (Applied Biosystems, CA, USA) thermocycler ine anotevera mamiriro kune mazhinji ekutanga mapeya: 94 °C kwemaminetsi mashanu, 5 kutenderera pa40 °C kwe94 s, 30 °C. ye58 s uye 45 °C ye72 s uye yekupedzisira elongation pa 45 °C kwemaminitsi mashanu. Kana iri ACM72 uye ACM5, kubata-pasi PCR kwakaiswa neannealing ye446 °C kusvika 449 °C pamusoro pematenderedzwa gumi, 60 kutenderera pa55 °C, ichiteverwa nekuwedzera kwekupedzisira kwemaminetsi mashanu pa10 °C. Zvigadzirwa zvePCR zvakaiswa mundiro ye30-tsime uye yakasanganiswa ne55 gL yeHi-Di Formamide (Life Technologies, Carlsbad, CA, USA) uye 5 gL GeneScan 72 ROX Size Standard (Life Technologies, Carlsbad, CA, USA). Amplicon yakagadziriswa neABI PRISM 96 Avant Genetic Analyzer (Life Technologies, Carlsbad, CA, USA) capillary sequencing muchina, uko alleles dzakapihwa seanotonga pamwe nekupihwa nekushandisa GeneMapper Software Version 14.
Iyo softwares GenAlEx 6.5 [57] uye Cervus 3.0.7 [58] akashandiswa kufungidzira nhamba yealleles (Na), nhamba yealleles inoshanda (Ne), akacherechedza heterozygosity (Ho), inotarisirwa heterozygosity (Iye), polymorphic information content (PIC), Shannon's information index (I), uye fixation index (Fis ) kune yega yega SSR locus.
Kuongororwa kweGenetic Diversity
Hierarchical partitioning of genetic variation pakati uye mukati mehanyanisi yaiongororwa neGenAlEx 6.5 [57] kuburikidza nekuongororwa kwema molecular variance (AMOVA) ine 999 bootstrapping kuti iongorore kukosha. Zvakare, GenAlEx 6.5 software yakashandiswa kufungidzira kusiyana-siyana mukati mehuwandu hwega hwega kuburikidza nekombuta avhareji yeHo, Iye, uye Fis pamusoro peSSR yese.
Mamiriro ehuwandu hwehuwandu hwehuwandu hwehuwandu hwehuwandu hwehuwandu hwehuwandu hwehuwandu hwakakonzerwa neBayesian model-based clustering algorithm inoshandiswa mu STRUCTURE v.2.3.4 software [59]. Iyo data set yaiitwa nehuwandu hwekufungidzira masumbu (K), kubva pa1 kusvika ku10, achiisa gumi akazvimirira kumhanya pane yega yega K kukosha. Pakumhanya kwega kwega, tichivavarira kuona kuenderana kwemhedzisiro, zviuru zana zvekutanga kupisa-munguva uye zana Markov Chain Monte Carlo (MCMC) iterations yakaitwa pasi peiyo admixture modhi uye yakazvimirira alle frequency frequency pakati pehuwandu. Iyo inonyanya kukosha K kukosha kwakatemwa kushandisa nzira yeAK, inotsanangurwa na [60], muchirongwa chewebhu-chakavakirwa STRUCTURE HARVESTER [61]. Huwandu hwevanhu hwakapihwa kune rimwe boka apo nhengo yaro coefficient (q-value) yaive yakakwira kupfuura 0.7, zvikasadaro yaionekwa seyakasanganiswa madzitateguru.
Principal coordination analysis yakaitwa kuitira kufungidzira mapatani ehukama hwemajini pakati pezviwanikwa zvakaratidzwa neNei's genetic distance matrix (Supplementary Table S5). Zvichienderana nemafambiro ezvese, dendrogram ye genetic distance yakagadzirwa kushandisa nzira isina uremu yeboka reviri ine arithmetic avhareji (UPGMA) cluster analysis muPOPTREEW software. [62]. Bootstrapping yakashandiswa kuongorora chivimbo mukusangana kwehierarchical, kuseta 100 resampling ye data set. Pakupedzisira, MEGA X software [63] yakashandiswa semuti wekudhirowa software.
Supplementary Materials: Izvi zvinotevera zviripo online http://www.mdpi.com/2223-7747/9/2/260/s1. Tafura S1: morphological maitiro eARO, MCO, uye TRO bulbs. Tafura S2: Heterozygosity uye fixation indices akaverengerwa ARO landraces uye TRO uye MCO landraces. Tafura S3: Pairwise kukosha kweiyo Fpt parameter. Tafura S4: Rondedzero yeSSR yakashandiswa muchidzidzo. Tafura S5. Pairwise population matrix yeNei genetic kureba. Mufananidzo S1: Chati yemutsara weK kukosha kuri kuchinja neEvanno's Delta K.
Munyori Mipiro: CL uye LR vakabata chidzidzo uye vakagadzira kuedza; CL uye PI yakaita molecular marker analysis; ARM neVZ vakaita bvunzo dzemunda; RM, SP, GR, uye CL zvakabatanidzwa mukuongorora data; RM uye CL vakanyora manyoro. Vese vanyori vakaverenga uye vakabvumirana neshanduro yakadhindwa yezvinyorwa.
Mariro: Iri basa rakatsigirwa nemari neRegional Apulian chirongwa "Biodiversity yeApulian miriwo yemarudzi" -Programma di Sviluppo Rurale per la Puglia 2014-2020. Misura 10—Sottomisura 10.2; kupa CUP H92C15000270002, Italy.
Kuonga: Kutenda kunobva kuna “Azienda Agricola Iannone Anna” uye “Associazione produttori della vera cipolla rossa di Acquaviva” nekupa zvinhu zvechirimwa zvakashandiswa mukuyedza.
Kurwisana kwekufarira: Vanyori vanoti hapana kupesana kwechido.
References
- 1. Stearn, WT Mangani marudzi eAllium anozivikanwa? Kew Mag. 1992, 9, 180-182. [CrossRef]
- 2. FAOSTAT. FAO Statistical Database. Inowanikwa online: http://www.fao.org/2017 (yakasvika musi wa8 Ndira 2019).
- 3. Block, E. The chemistry yegariki uye hanyanisi. Sci. Am. 1985, 252, 114-119. [CrossRef]
- 4. Lee, B.; Jung, JH; Kim, HS Ongororo yehanyanisi tsvuku pane antioxidant chiitiko mumakonzo. Food Chem. Toxicol. 2012, 50, 3912-3919. [CrossRef]
- 5. Lee, SM; Mwedzi, J.; Chung, JH; Cha, YJ; Shin, MJ Mhedzisiro ye quercetin-inopfuma yehanyanisi peel inotorwa pane arterial thrombosis mumakonzo. Food Chem. Toxicol. 2013, 57, 99-105. [CrossRef] [PubMed]
- 6. Yoshinari, O.; Shiojima, Y.; Igarashi, K. Anti-obesity zvinokonzerwa neonion extract muzucker diabetic fatty rats. Nutrients 2012, 4,1518-1526. [CrossRef]
- 7. Akash, MSH; Rehman, K.; Chen, S. Spice chirimwa Allium cepa: Dietary supplement yekurapa rudzi rwe 2 chirwere cheshuga mellitus. udyo 2014, 30, 1128-1137. [CrossRef] [PubMed]
- 8. Wang, Y.; Tian, WX; Ma, XF Inhibitory Migumisiro yehanyanisi (Allium cepa L.) inobvisa pakuwanda kwekenza maseru uye adipocytes kuburikidza nekudzivisa fatty acid synthase. Asian Pac. J. Cancer Prev. 2012,13, 5573-5579. [CrossRef] [PubMed]
- 9. Lai, WW; Hsu, SC; Chueh, FS; Chen, YY; Yang, JS; Lin, JP; Lien, JC; Tsai, CH; Chung, JG Quercetin inhibits kutama uye kupinda kweSAS masero emukenza wemuromo wevanhu kuburikidza nekudzivisa NF-kappaB uye matrix metalloproteinase-2 / -9 nzira dzechiratidzo. Anticancer Res. 2013, 33, 1941-1950. [PubMed]
- 10. Nicastro, HL; Ross, SA; Milner, JA Garlic uye hanyanisi: Zvivakwa zvavo zvekudzivirira gomarara. Cancer Prev. Res. 2015, 8,181-189. [CrossRef]
- 11. Forte, L.; Torricelli, P.; Boanini, E.; Gazzano, M.; Rubini, K.; Fini, M.; Bigi, A. Antioxidant uye bone kugadzirisa zvinhu zvequercetin-functionalized hydroxyapatite: An in vitro osteoblast-osteoclast-endothelial cell co-culture study. Acta Biomater. 2016, 32, 298-308. [CrossRef]
- 12. Yamazaki, Y.; Iwasaki, K.; Mikami, M.; Yagihashi, A. Distribution of eleven flavour precursors, S-Alk(en)yl-L-cysteine derivatives, mumiriwo minomwe yeAllium. Food Sci. Technology. Res. 2011, 17, 55-62. [CrossRef]
- 13. Block, E. The organosulphur chemistry of the Genus Allium—Zvinorehwa neorganic chemistry yesarufa. Angew. Chem. Int. Ed. Chirungu. 1992, 31, 1135-1178. [CrossRef]
- 14. Griffiths, G.; Trueman, L.; Crowther, T.; Thomas, B.; Smith, B. Hanyanisi-Kubatsira kwepasi rose kune hutano. Phytother. Res. 2002,16, 603-615. [CrossRef]
- 15. Schwimmer, S.; Weston, WJ Enzymatic kukura kwepyruvic acid muanisi seyero yepungency. J. Agric. Food Chem. 1961, 9, 301-304. [CrossRef]
- 16. Ketter, CAT; Randle, WM Pungency ongororo muhanyanisi. In Yakaedzwa Zvidzidzo zveLabhoretari Kudzidzisa; Karcher, SJ, Ed.; Association for Biology Laboratory Education (ABLE): New York, NY, USA, 1998; Vhoriyamu 19, mapeji 177-196.
- 17. Hanelt, P Taxonomy, evolution, uye nhoroondo. In Hanyanisi neChirimwa Chakabatana, Vol. I. Botany, Physiology uye Genetics; Rabinowitch, HD, Brewster, JL, Eds.; CRC Press: Boca Raton, FL, USA, 1990; map. 1-26.
- 18. Rabinowitch, HD; Kura, L. Allium Crop Science: Recent Advances; CABI Publishing: Wallingford, UK, 2002.
- 19. Mallor, C.; Carravedo, M.; Estopanan, G.; Mallor, F. Hunhu hwemajini zviwanikwa zveanisi (Allium cepa L.) kubva kuSpanish sekondari nzvimbo yekusiyana. Span. J. Agric. Res. 2011, 9, 144-155. [CrossRef]
- 20. Ferioli, F.; D'Antuono, LF Kuongororwa kwe phenolics uye cysteine sulfoxides muanisi yemunharaunda uye shallot germplasm kubva kuItaly neUkraine. Genet. Resour. Crop Evol. 2016, 63, 601-614. [CrossRef]
- 21. Petropoulos, SA; Fernandes, A.; Barros, L.; Ferreira, ICFR; Ntatsi, G. Morphological, nutritional and chemical description of 'vatikiotiko', anion local landrace kubva kuGreece. Food Chem. 2015,182, 156-163. [CrossRef]
- 22. Liguori, L.; Adiletta, G.; Nazzaro, F.; Fratianni, F.; Di Matteo, M.; Albanese, D. Biochemical, antioxidant properties uye antimicrobial chiitiko chemhando dzakasiyana dzehanyanisi munzvimbo yeMediterranean. J. Food Meas. Hunhu. 2019,13, 1232-1241. [CrossRef]
- 23. Yoo, KS; Pike, L.; Crosby, K.; Jones, R.; Leskovar, D. Musiyano wehanyanisi pungency nekuda kwezvirimwa, nharaunda yekukura, uye saizi yegirobhu. Sci. Hortic. 2006,110, 144-149. [CrossRef]
- 24. Beesk, N.; Perner, H.; Schwarz, D.; George, E.; Kroh, LW; Rohn, S. Kugoverwa kwe quercetin-3, 4'-O-diglucoside, quercetin-4'-O-monoglucoside, uye quercetin muzvikamu zvakasiyana zvegirobhu rehanyanisi (Allium cepa L.) inokonzerwa negenotype. Food Chem. 2010,122, 566-571. [CrossRef]
- 25. Caruso, G.; Conti, S.; Villari, G.; Borrelli, C.; Melchionna, G.; Minutolo, M.; Russo, G.; Amalfitano, C. Mhedzisiro yekudyara nguva uye kusimba kwechirimwa pagoho, kunaka uye antioxidant zviri mukati mehanyanisi. (Allium cepa L.) kumaodzanyemba kweItaly. Sci. Hortic. 2014,166, 111-120. [CrossRef]
- 26. Perez-Gregorio, MR; Regueiro, J.; Simal-Gandara, J.; Rodrigues, AS; Almeida, DPF Kuwedzera iyo yakawedzera-kukosha kwehanyanisi sesosi ye antioxidant flavonoids: Ongororo yakakosha. Crit. Rev. Food Sci. Nutriti. 2014, 54,1050-1062. [CrossRef] [PubMed]
- 27. Pohnl, T.; Schweiggert, RM; Carle, R. Mhedzisiro yenzira yekurima uye kusarudzwa kwecultivar pane soluble carbohydrates uye pungent misimboti muhanyanisi. (Allium cepa L.). J. Agric. Food Chem. 2018, 66, 12827-12835. [CrossRef] [PubMed]
- 28. Tedesco, I.; Carbone, V.; Spagnuolo, C.; Minasi, P.; Russo, GL Kuzivikanwa uye quantification yeflavonoids kubva kumarudzi maviri ekumaodzanyemba eItaly cultivars. allium cepa L., Tropea (red onion) uye Montoro (copper onion), uye kukwanisa kwavo kuchengetedza erythrocytes yevanhu kubva kune oxidative stress. J. Agric. Food Chem. 2015, 63, 5229-5238. [CrossRef]
- 29. Villano, C.; Esposito, S.; Carucci, F.; Frusciante, L.; Carputo, D.; Aversano, R. High-throughput genotyping in onion inoratidza chimiro chemajini akasiyana uye anodzidzisa SNPs anobatsira pakurera mamorekuru. Mol. Breed. 2019, 39, 5. [CrossRef]
- 30. Mercati, F.; Longo, C.; Poma, D.; Araniti, F.; Lupini, A.; Mammano, MM; Fiore, MC; Abenavoli, MR; Sunseri, F Genetic mutsauko weItaly refu sherufu-upenyu madomasi (Solanum lycopersicum L.) muunganidzwa uchishandisa SSR uye morphological michero hunhu. Genet. Resour. Crop Evol. 2014, 62, 721-732. [CrossRef]
- 31. Gonzalez-Perez, S.; Mallor, C.; Garces-Claver, A.; Merino, F.; Taboada, A.; Rivera, A.; Pomar, F.; Perovic, D.; Silvar, C. Kuongorora genetic diversity uye hunhu hwemhando muunganidzwa wehanyanisi (Allium cepa L.) nzvimbo kubva kuchamhembe-kumadokero kweSpain. Genetics 2015, 47, 885-900. [CrossRef]
- 32. Roti, C.; Iovieno, P.; Centomani, I.; Marcotrigiano, AR; Fanelli, V.; Mimiola, G.; Summo, C.; Pavan, S.; Ricciardi, L. Genetic, bio-agronomic, uye kudya kunovaka kwekare (brassica oleracea L. var. acephala) kusiyana-siyana muApulia, Southern Italy. Diversity 2018,10, 25. [CrossRef]
- 33. Bardaro, N.; Marcotrigiano, AR; Bracuto, V.; Mazzeo, R.; Ricciardi, F.; Loti, C.; Pavan, S.; Ricciardi, L. Genetic kuongororwa kwekupikisa Orobanche crenata (Forsk.) mupea (Pisum sativum L.) mutsara wakaderera-strigolactone. J. Plant Pathol. 2016, 98, 671-675.
- 34. Wako, T.; Tsukazaki, H.; Yaguchi, S.; Yamashita, K.; Ito, S.; Shigyo, M. Mepu yehuwandu hwemaitiro loci yenguva yekubolting mubunching hanyanisi (Allium fistulosum L.). Euphytica 2016, 209, 537-546. [CrossRef]
- 35. Dhaka, N.; Mukhopadhyay, A.; Paritosh, K.; Gupta, V.; Pental, D.; Pradhan, AK Kuzivikanwa kweiyo genic SSRs uye kuvakwa kweSSR-yakavakirwa mepu yekubatanidza mukati Brassica juncea. Euphytica 2017, 213, 15. [CrossRef]
- 36. Anandhan, S.; Mote, SR; Gopal, J. Kuongorora kweonion varietal identity uchishandisa zviratidzo zveSSR. Mbeu Sci. Technology. 2014, 42, 279-285. [CrossRef]
- 37. Mitrova, K.; Svoboda, P.; Ovesna, J. Kusarudzwa uye kusimbiswa kwechiratidzo seti yekusiyanisa mbeu dzehanyanisi kubva kuCzech Republic. Czech J. Genet. Plant Breed. 2015, 51, 62-67. [CrossRef]
- 38. Di Rienzo, V.; Miazzi, MM; Fanelli, V.; Sabetta, W.; Montemurro, C. Kuchengetedzwa uye hunhu hweApulian olive germplasm biodiversity. Acta Hortic. 2018,1199,1-6. [CrossRef]
- 39. Mallor, C.; Arnedo-Andres, A.; Garces-Claver, A. Kuongorora iyo genetic kusiyana kweSpanish allium cepa landraces yekupfuya hanyanisi uchishandisa ma microsatellite marker. Sci. Hortic. 2014,170, 24-31. [CrossRef]
- 40. Rivera, A.; Mallor, C.; Garces-Claver, A.; Garcia-Ulloa, A.; Pomar, F.; Silvar, C. Kuongorora genetic kusiyana muhanyanisi (allium cepa L.) landraces kubva kuchamhembe kwakadziva kumadokero kweSpain uye kuenzanisa nekusiyana kweEurope. NZJ Crop Hortic. 2016, 44, 103-120. [CrossRef]
- 41. De Giovanni, C.; Pavan, S.; Taranto, F.; Di Rienzo, V.; Miazzi, MM; Marcotrigiano, AR; Mangini, G.; Montemurro, C.; Ricciardi, L.; Lotti, C. Genetic variation of a global germplasm collection of chickpea (Cicer arietinum L.) kusanganisira kupinda kweItaly panjodzi yekukukurwa kwemajini. Physiol. Mol. Biol. Zvirimwa 2017, 23, 197-205. [CrossRef]
- 42. Mazzeo, R.; Morgese, A.; Sonnante, G.; Zuluaga, DL; Pavan, S.; Ricciardi, L.; Lotti, C. Genetic diversity mu broccoli rabe (Brassica rapa L. subsp. sylvestris (L.) Janch.) kubva kuSouth Italy. Sci. Hortic. 2019, 253, 140-146. [CrossRef]
- 43 Jakse, M.; Martin, W.; McCallum, J.; Havey, M. Single nucleotide polymorphisms, indels, uye nyore kutevedzana kunodzokororwa kweiyo onion cultivar identification. J. Am. Soc. Hortic. Sci. 2005,130, 912-917. [CrossRef]
- 44. McCallum, J.; Thomson, S.; Pither-Joyce, M.; Kenel, F. Genetic diversity analysis uye single-nucleotide polymorphism marker development mugirobhu rinorimwa hanyanisi zvichibva pakutevedzana kwetag-nyore kutevedzana kudzokorora mamakisi. J. Am. Soc. Hortic. Sci. 2008,133, 810-818. [CrossRef]
- 45. Baldwin, S.; Pither-Joyce, M.; Wright, K.; Chen, L.; McCallum, J. Kuvandudzwa kweiyo robust genomic yakapusa sequence inodzokorora mamakisi yefungidziro yemajini akasiyana mukati uye pakati pegirobhu hanyanisi. (Allium cepa L.) vanhu. Mol. Breed. 2012, 30, 1401-1411. [CrossRef]
- 46. DeWoody, JA; Honeycutt, RL; Skow, LC Microsatellite mamaki mune chena muswe deer. J. Hered. 1995, 86, 317-319. [CrossRef] [PubMed]
- 47. Khodadadi, M.; Hassanpanah, D. Iranian hanyanisi (Allium cepa L.) cultivars mhinduro kune inbreeding depression. World Appl. Sci. J. 2010,11, 426-428.
- 48. Abdou, R.; Bakasso, Y.; Saadou, M.; Baudoin, JP; Hardy, OJ Genetic kusiyana kweNiger hanyanisi (Allium cepa L.) yakaongororwa neakapusa kutevedzana kudzokorora mamaki (SSR). Acta Hortic. 2016,1143, 77-90. [CrossRef]
- 49. Pavan, S.; Loti, C.; Marcotrigiano, AR; Mazzeo, R.; Bardaro, N.; Bracuto, V.; Ricciardi, F.; Taranto, F.; D'Agostino, N.; Schiavulli, A.; et al. Iyo yakasarudzika genetic cluster mune yakarimwa chickpea sezvakaratidzwa negenome-wide marker kuwanikwa uye genotyping. Plant Genome 2017, 2017,10. [CrossRef]
- 50. Pavan, S.; Marcotrigiano, AR; Ciani, E.; Mazzeo, R.; Zonno, V.; Ruggieri, V.; Loti, C.; Ricciardi, L. Genotyping-nekutevedzana kwevise (Cucumis melo L.) Kuunganidzwa kwegermplasm kubva kune yechipiri nzvimbo yekusiyana-siyana kunosimbisa mapatani ekusiyana kwemajini uye genomic maficha emhando dzakasiyana madziva. BMC Genom. 2017, 18, 59. [CrossRef]
- 51. Di Rienzo, V.; Zion, S.; Taranto, F.; D'Agostino, N.; Montemurro, C.; Fanelli, V.; Sabetta, W.; Boucheffa, S.; Tamendjari, A.; Pasqualone, A.; et al. Genetic inoyerera pakati pehuwandu hwemiorivhi mukati meMediterranean basin. Peer J. 2018, 6. [CrossRef]
- 52. Mufudzi, LD; McLay, TG Maviri madiki-scale mapuroteni ekuparadzaniswa kweDNA kubva kune polysaccharide-rich chirimwa tishu. J. Plant Res. 2011,124, 311-314. [CrossRef]
- 53. Doyle, JJ; Doyle, JL Kusarudzika kwechirimwa DNA kubva munyama nyowani. Focus 1990,12, 13-14.
- 54 Kuhl, JC; Cheung, F.; Qiaoping, Y.; Martin, W.; Zewdie, Y.; McCallum, J.; Catanach, A.; Rutherford, P.; Sink, KC; Jenderek, M.; et al. Imwe yakasarudzika seti ye11,008 hanyanisi yakaratidza kutevedzana ma tag inoratidza yakaratidza kutevedzana uye genomic kusiyana pakati pe monocot mirairo asparagales uye poales. Dyara Sero 2004,16, 114-125. [CrossRef]
- 55. Kim, HJ; Lee, HR; Hyun, JY; Song, KH; Kim, KH; Kim, JE; Hur, CG; Harn, CH Mucherechedzo kuvandudzwa kweonion genetic kuchena yekuongorora uchishandisa SSR Finder. Korean J. Breed. Sci. 2012, 44, 421-432. [CrossRef]
- 56. Schuelke, M. Nzira yehupfumi yekunyorwa kwefluorescent yePCR zvidimbu. Nat. Biotechnol. 2000, 18, 233-234. [CrossRef] [PubMed]
- 57 Peakall, R.; Smouse, PE GenAlEx 6.5: Genetic analysis muExcel. Population genetic software yekudzidzisa nekutsvaga: An update. Bioinformatics 2012, 28, 2537-2539. [CrossRef] [PubMed]
- 58. Kalinowski, ST; Taper, ML; Marshall, TC Kuongorora kuti chirongwa chekombuta CERVUS chinogadzika sei genotyping kukanganisa kunowedzera budiriro mukupihwa kwababa. Mol. Ecol. 2007,16, 1099-1106. [CrossRef]
- 59. Pritchard, JK; Stephens, M.; Rosenberg, NA; Donnelly, P. Association mepu muhuwandu hwakarongeka. Am. J. Hum. Genet. 2000, 67, 170-181. [CrossRef]
- 60. Evanno, G.; Regnaut, S.; Goudet, J. Kuona nhamba yemasumbu evanhu vanoshandisa software STRUCTURE: Chidzidzo chekufananidza. Mol. Ecol. 2005,14, 2611-2620. [CrossRef]
- 61. Earl, D.; VonHoldt, B. STRUCTURE HARVESTER: Webhusaiti uye chirongwa chekuona STRUCTURE kubuda uye kushandisa nzira yeEvanno. Conserv. Genet. Resour. 2011, 4. [CrossRef]
- 62. Takezaki, N.; Nei, M.; Tamura, K. POPTREEW: Webhu vhezheni yePOPTREE yekuvaka miti yehuwandu kubva kune alle frequency data uye komputa humwe humwe huwandu. Mol. Biol. Evol. 2014, 31, 1622-1624. [CrossRef]
- 63. Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X. Molecular Evolutionary Genetics Analysis pamapuratifomu emakomputa. Mol. Biol. Evol. 2018, 35, 1547-1549. [CrossRef]